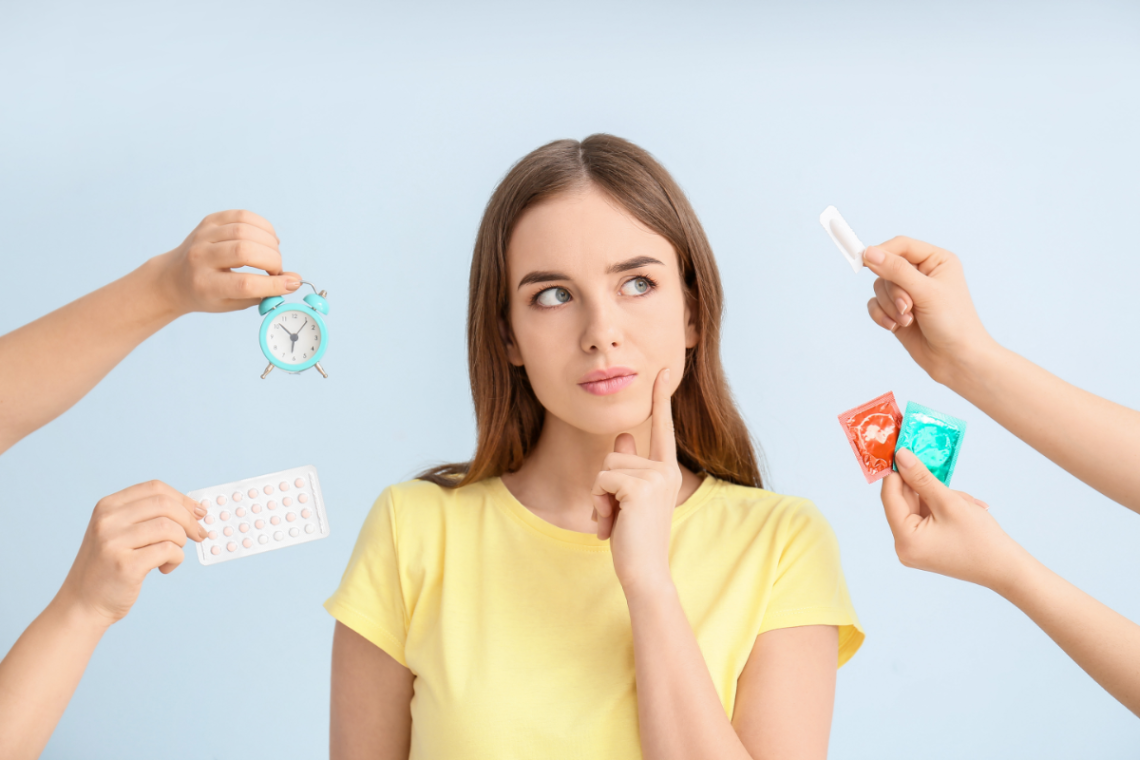
Les alternatives à la pilule contraceptive: opter pour une contraception naturelle et non hormonale
Dans ce monde moderne, il existe plusieurs options pour les femmes qui souhaitent contrôler leur fertilité et pratiquer une planification familiale sans recourir aux hormones synthétiques. Les contraceptifs hormonaux tels que la pilule, l’implant et l’anneau vaginal sont très populaires en raison de leur efficacité. Cependant, de plus en plus de femmes se tournent vers des méthodes non hormonales telles que le stérilet au cuivre, le diaphragme et la cape cervicale pour éviter les effets secondaires et les problèmes de santé liés à ces hormones artificielles. L’intérêt croissant pour les méthodes contraceptives naturelles Certaines femmes rencontrent des difficultés avec les…
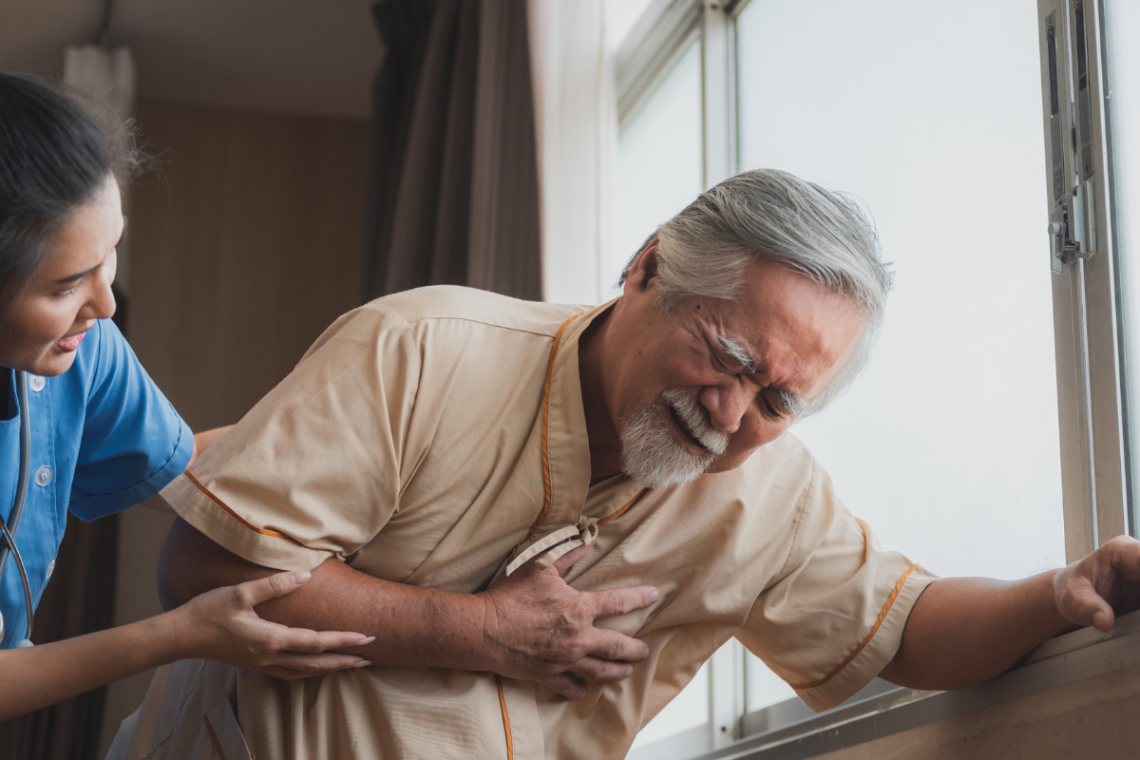
L’importance de l’ECG dans le suivi des patients atteints de troubles du rythme cardiaque
Les troubles du rythme cardiaque sont une préoccupation majeure pour les professionnels de la santé, car ils peuvent provoquer des complications graves si elles ne sont pas surveillées et gérées efficacement. L’électrocardiogramme (ECG) est un outil essentiel pour détecter et suivre les arythmies cardiaques chez les patients. Dans cet article, nous examinerons comment l’ECG aide à la gestion des patients atteints de troubles du rythme cardiaque. Le rôle de l’électrocardiogramme (ECG) Un ECG est un test non invasif qui enregistre les séquences électriques des battements cardiaques. Il s’agit d’une méthode précise et rapide pour évaluer la fonction cardiaque, identifier toute…
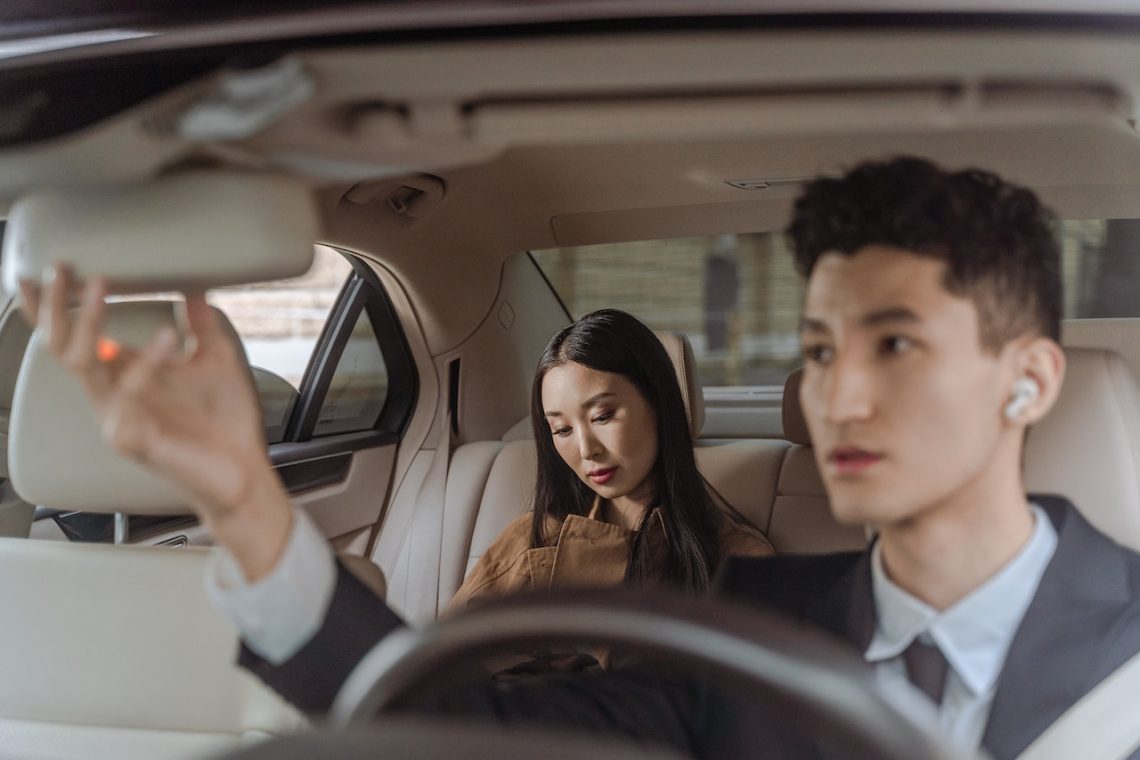
Améliorez votre expérience lors de la conduite sur longue distance grâce aux coussins de voiture
Les trajets en voiture peuvent parfois être inconfortables, surtout lorsque vous devez parcourir de longues distances. Les coussins de voiture sont une excellente solution pour améliorer le confort et l’expérience de conduite. Dans cet article, nous allons aborder l’utilité des coussins de voiture, les différents types disponibles et comment choisir celui qui convient le mieux à vos besoins pour une conduite agréable. Les avantages des coussins de voiture Utiliser un coussin pour la voiture offre plusieurs avantages : Réduction des douleurs et tensions musculaires : Un bon coussin de voiture permet de maintenir une position ergonomique et de réduire les…
7 conseils pour renforcer votre confiance en soi à travers l’hypnose et le développement personnel
La confiance en soi est un élément essentiel pour réussir dans la vie, tant sur le plan professionnel que personnel. Elle permet d’affronter les difficultés avec assurance et de s’adapter aux changements avec aisance. Découvrez ces 7 conseils pour booster votre confiance en soi grâce à des techniques telles que l’hypnose, le renforcement de l’estime de soi et les ateliers de développement personnel. 1. Identifier les sources de votre manque de confiance Pour renforcer votre confiance en vous, il faut d’abord comprendre ce qui vous empêche de croire en vous-même. Prenez le temps d’analyser vos pensées et vos comportements afin…
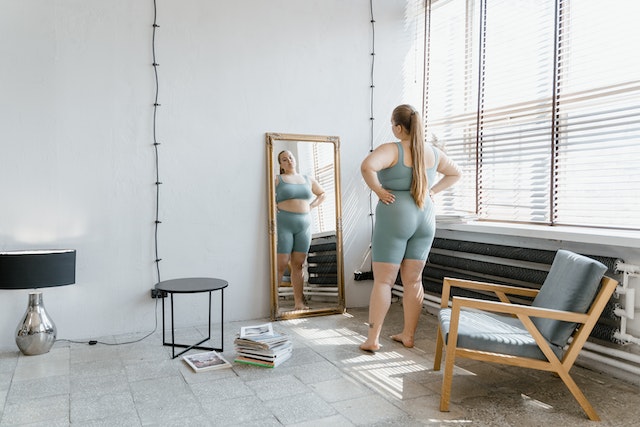
IMC et santé : comment utiliser l’indice de masse corporelle pour évaluer votre état de santé général ?
L’indice de masse corporelle (IMC) est un critère de santé utilisé pour évaluer la composition corporelle d’une personne et déterminer le poids idéal. Il est essentiel de calculer son IMC pour comprendre l’état de sa santé et prévenir les troubles liés à l’obésité et à la dénutrition, mais aussi pour obtenir une assurance ou un prêt immobilier. Le calcul de l’IMC se base sur la taille et le poids d’une personne, et permet de repérer rapidement toute anomalie qui nécessite un suivi médical. La prise en compte de ces repères permet ainsi de mieux comprendre son état de santé et…
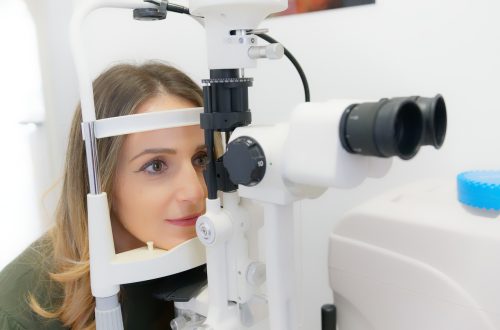
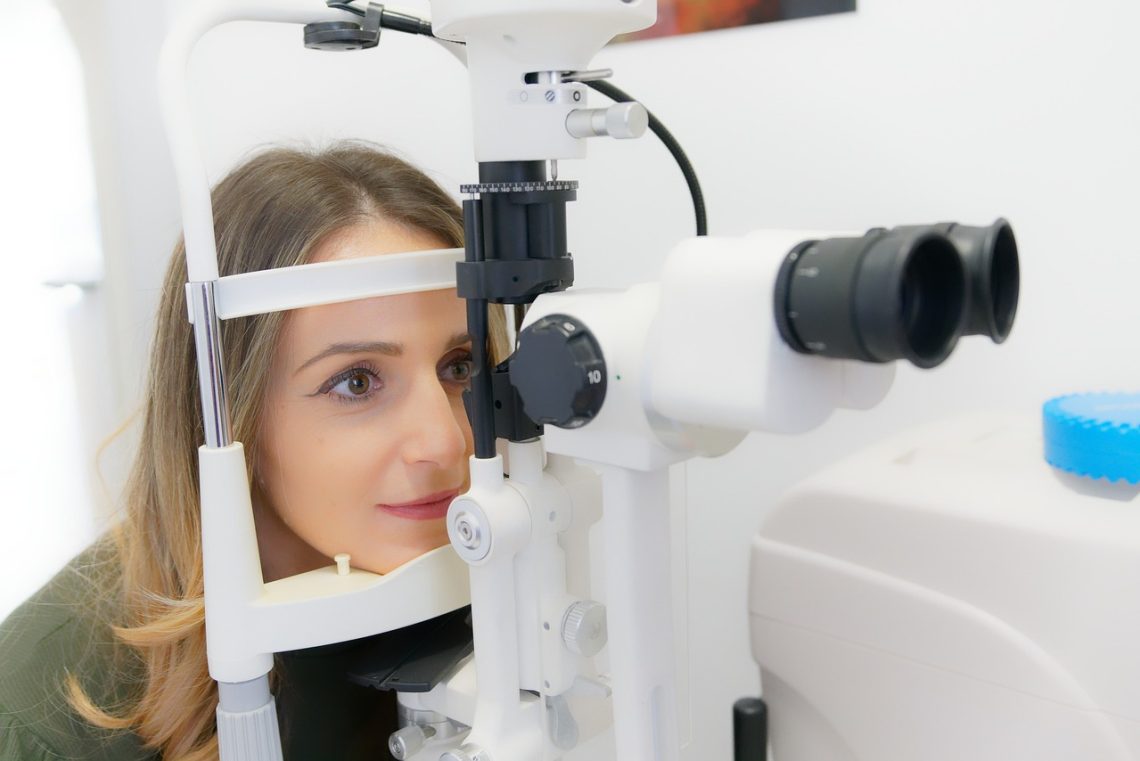
Que savoir de l’opération laser des yeux ?
Vous ne le saviez peut-être pas, mais vos yeux peuvent bel et bien subir une opération chirurgicale en vue de les rendre plus performants. En réalité, ces organes sont très sensibles et s’exposent à plusieurs anomalies visuelles telles que la myopie, l’hypermétropie, etc. Heureusement, il y a moyen d’y remédier avec une chirurgie au laser des yeux. Mais avant, gros plan sur cette forme d’intervention et sur ce qu’il importe d’en retenir avant d’y prétendre. Chirurgie des yeux au laser, de quoi s’agit-il ? La chirurgie au laser des yeux n’est rien d’autre qu’une intervention particulièrement orientée vers les yeux et…
Comment prévenir les troubles musculo-squelettiques sur le lieu de travail ?
Garder vos employés en bonne santé et productifs est essentiel pour la réussite de votre entreprise. Cependant, ce n’est pas toujours facile à réaliser. Si vous travaillez dans un environnement de travail manuel ou si votre emploi exige de rester assis ou debout pendant de longues heures, vous pouvez avoir du mal à assurer la sécurité et la santé de vos employés au quotidien. Le fait de travailler dans un bureau peut également être facile à comprendre pour certains d’entre vous. Avec l’augmentation de la prévalence des emplois de bureau, les gens ont tendance à rester assis pendant la majorité…
Comment choisir une mutuelle pour seniors avec un bon remboursement sur l’optique ?
Lorsque vous atteignez 65 ans, votre vie change à plus d’un titre. Votre avenir change parce que vous devez maintenant penser à votre retraite et à la façon dont vous allez la financer. Votre présent change parce que votre santé et votre vie sociale deviennent plus importantes. Même votre passé change ; c’est le moment où vous devenez éligible à des prestations pour personnes âgées telles que des services médicaux à prix réduit et d’autres avantages conçus pour aider les personnes âgées à mener une vie heureuse et saine. Pour vous aider à tirer le meilleur parti de vos années…
Comment trouver un médecin généraliste sans rendez-vous à Bobigny
Les services des cliniques sans rendez-vous ne sont pas réservés à la saison de la grippe ou aux entorses occasionnelles à la cheville. Si vous vivez dans une ville animée comme Bobigny, il peut être difficile d’obtenir un rendez-vous chez votre médecin traitant pendant les heures d’ouverture normales. Si vous avez besoin d’une attention médicale urgente et que vous avez épuisé toutes les autres options, vous pouvez vous rendre dans une clinique sans rendez-vous. Ces cliniques sont conçues pour fournir des soins rapides et abordables pour les maladies et blessures mineures, avec un temps d’attente minimal. Trouver un médecin généraliste…
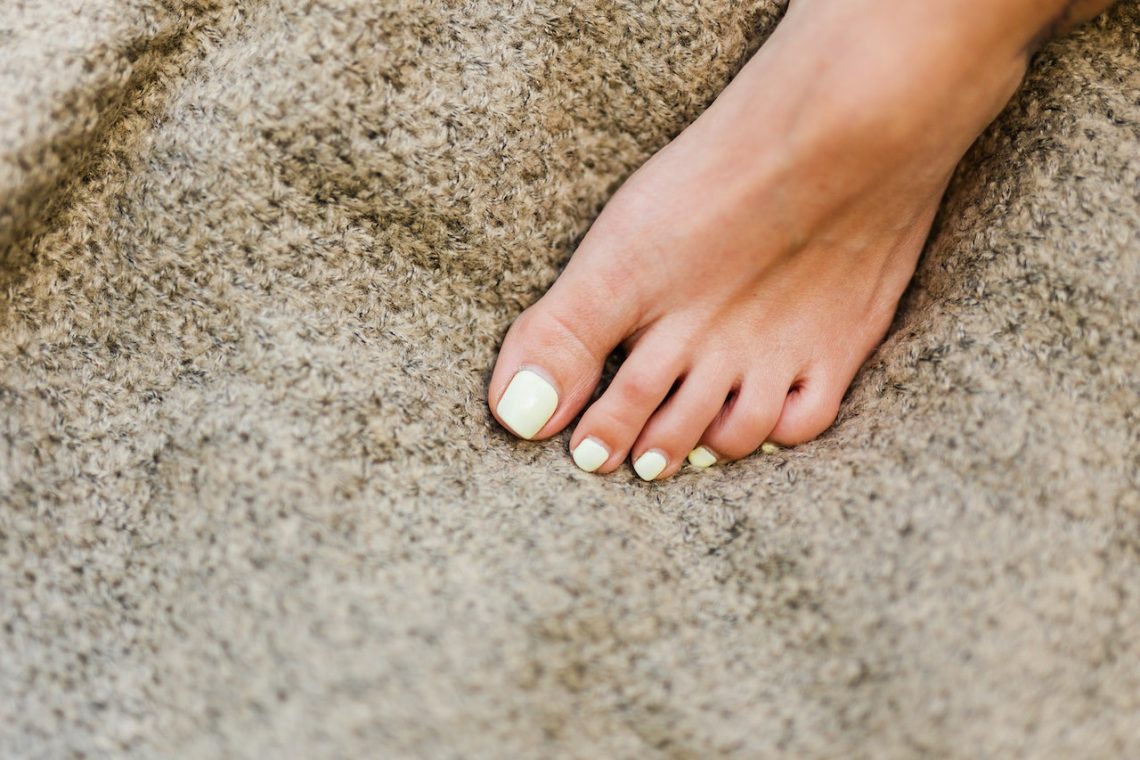
Temps de guérison d’un orteil cassé : 5 conseils pour vous aider à guérir rapidement
Toute fracture de l’orteil ou du pied peut être douloureuse, mais une fracture du cinquième métatarsien est particulièrement gênante. Appelé l’avant-pied parce qu’il s’agit de l’os le plus grand, le cinquième métatarsien est sujet aux fractures en raison de son emplacement et prend donc plus de temps que les autres os du pied pour guérir. C’est également l’un des endroits les plus courants où les personnes ayant des pieds plats souffrent de fractures. La bonne nouvelle est que si vous prenez soin de vous et suivez quelques conseils pour une guérison plus rapide, vous pourrez vous sentir mieux et vous…
Comment enlever la cire de ses cheveux ?
L’utilisation de la cire pour cheveux offre plusieurs avantages à son utilisateur. Toutefois, il est important de retirer ce produit capillaire avant d’aller au lit. Bien que cet exercice soit difficile à réaliser, il est cependant important de le faire pour éviter certains problèmes. Voici quelques astuces qui vous permettront de retirer la cire de vos cheveux en un laps de temps. Utiliser un shampoing pour retirer la cire de vos cheveux Si vous désirez avoir une excellente coupe de coiffure, vous pouvez utiliser la cire Red one. Cette cire pour homme s’adapte à tout type de cheveux et permet…
La pizza surgelée est-elle vraiment si saine ?
Nous savons tous que la pizza surgelée n’est pas le repas le plus sain qui soit. Mais parfois, après une longue journée de travail, la dernière chose que nous voulons faire est de préparer un dîner compliqué. Nous nous tournons donc vers la pizza surgelée comme option rapide et facile. Mais est-ce vraiment si sain ? Regardons cela de plus près. Des pizzas surgelées de plus en plus qualitatives La pizza surgelée a fait du chemin ces dernières années. Dans le passé, les pizzas surgelées étaient chargées de sodium et de graisses malsaines. Mais aujourd’hui, de nombreuses marques proposent des…
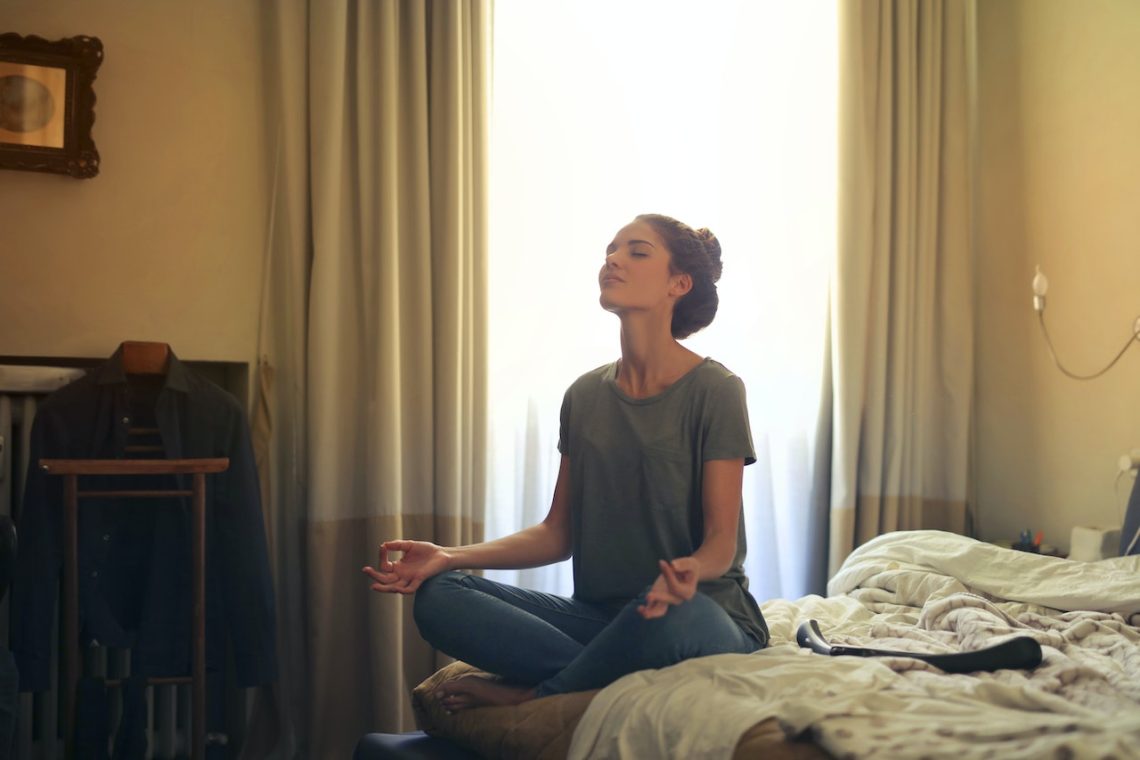
7 bienfaits de la méditation
Pour créer le calme dans votre esprit et réduire certaines douleurs, la méditation est une solution adéquate. Cette pratique inspirée du bouddhisme est de plus en plus adoptée par de nombreuses personnes. En effet, si la méditation connait un si grand succès, c’est à cause de ses effets positifs qui permettent d’atteindre un état de bien-être. Selon certains adeptes et professionnels du domaine, cela aurait de nombreux bienfaits sur le plan psychique comme sanitaire. Voici 7 bienfaits de la méditation ! Réduction du stress et de la dépression Plusieurs études ont révélé que pratiquer la méditation au quotidien avait des effets…
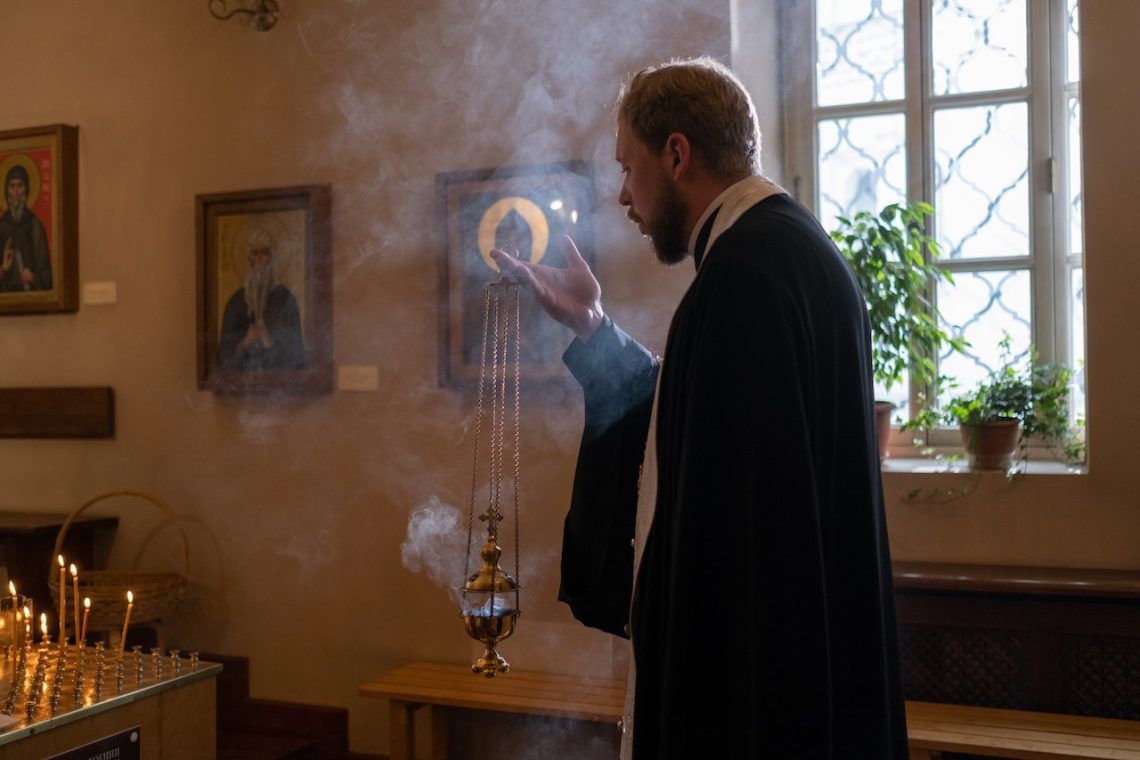
2 modèles d’encensoirs
Pouvoir vivre dans un environnement sain et rempli de toutes les ondes positives est ce que souhaite la plupart des gens. Pour ce faire, il faut prendre certaines habitudes comme purifier sa maison, son bureau ou encore son magasin. C’est ce à quoi l’encensoir vous aide. Pensez à opter pour les modèles que voici. L’encensoir électrique Cet accessoire est un objet millénaire qui a connu une certaine évolution dans le temps. Auparavant, il était plus connu sous la désignation bruleur d’encens. Aujourd’hui, le nom, mais aussi le design a évolué. L’encensoir électrique est le parfait exemple de la modernité qu’a…
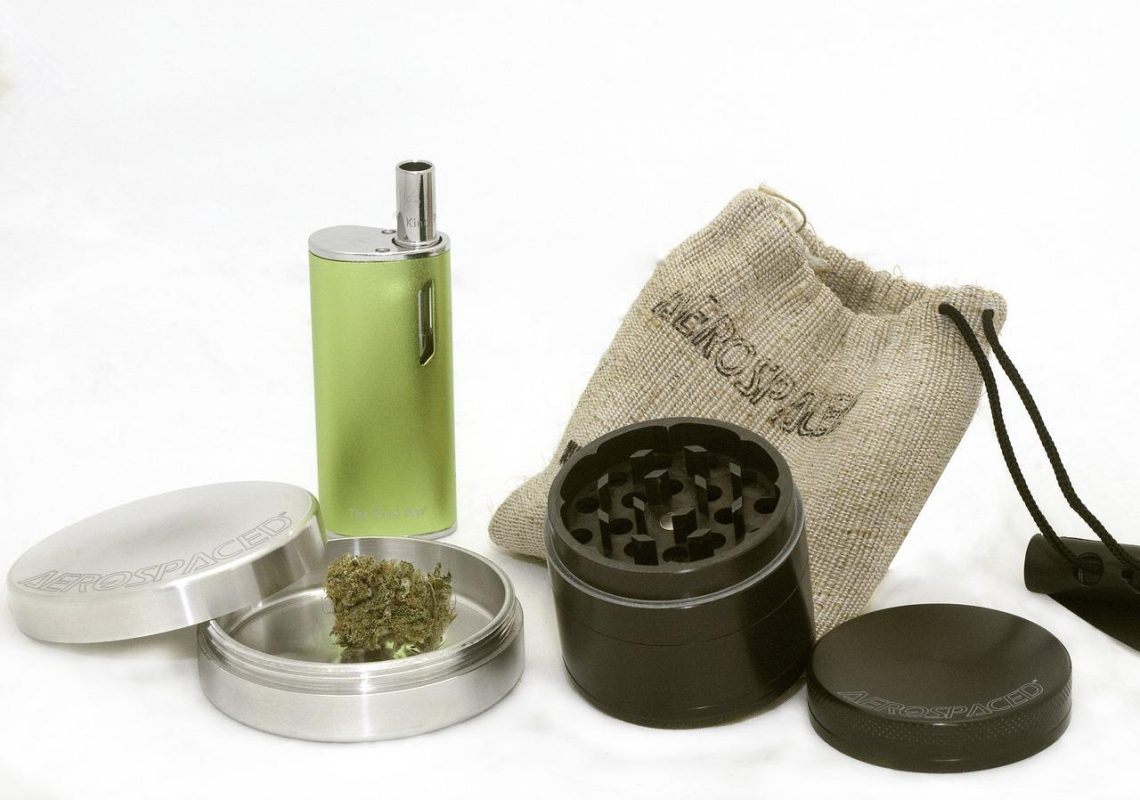
Protéger sa weed : quelles sont les alternatives au pocheton ?
La marijuana existe sous forme d’herbe et la couche juvénile de la société l’a surnommé la weed. Cette dernière procure des effets de détente et de bien-être au fumeur. Cependant, sa qualité pourrait être affectée si elle n’est pas correctement conservée. Le pocheton à weed est l’accessoire par excellence pour assurer cette conservation, mais il existe d’autres alternatives que vous découvrirez dans cet article. Pochon de weed Un amateur du cannabis a généralement un pocheton pour protéger sa weed de l’humidité et du vent. Ce gadget est effectivement protecteur et il permet de garder la weed en bonne qualité. Le…